When I run GESS I get the following warning: /Library/Frameworks/amework/Versions/2.7/lib/python2.7/site-packages/Bio/Seq.py:341: BiopythonDeprecationWarning: This method is obsolete please use str(my_seq) instead of my_seq.tostring().Īnd then I get the following error after Constructing SpliceGraph: Exon gap linking.: "Traceback (most recent call last):įile "/Library/Frameworks/amework/Versions/2.7/lib/python2.7/site-packages/gess/GESS.py", line 122, in įile "/Library/Frameworks/amework/Versions/2.7/lib/python2.7/site-packages/gess/GESS.py", line 64, in main I used JAGuaR (Junction Alignments to Genome for RNA-Seq Reads) for alignment. # Bayes factors thresholds to use for -plot-bf-dist # Bar color for Bayes factor distribution # (Used to normalize the read density for RPKM calculation) # List of colors for read denisites of each sample # Whether to plot the number of reads in each junction # Whether to plot posterior distributions inferred by MISO # Whether to use a log scale or not when plotting # Factor to scale down introns and exons by Miso_files = Dimensions of figure to be plotted (in inches) Miso_prefix = /home/ubuntu/SAMPLE1/miso_out Saving plot to: is the entire sashimi_plot_settings.txt file that with the changes I made: Hello I edited the sashimi_plot_settings.txt file but I still get this error Warning: MISO file not found 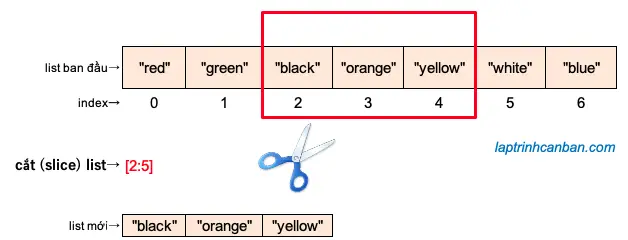
Please let me know if you need any further help. I hope this will help you finding the alternative splicing events. sashimi_plot_settings.txt -output-dir plots/
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |


 RSS Feed
RSS Feed
